I expect nearly every creationist and Intelligent Design blog will eventually discuss this, but I thought I would throw in my “two cents” about a study that has serious implications for the creation/evolution controversy. Laura Poliseno and her colleagues have published a study in Nature that has demonstrated a function for a class of pseudogenes.1 The study will result in a radical change in biology’s understanding of what has been disparagingly called “junk DNA.”
Let’s start from the beginning. A pseudogene is a section of DNA that looks a lot like a gene that exists in another section of an organism’s genome. However, despite this similarity, the pseudogene does not produce a protein. In other words, suppose a researcher finds a gene that produces a given protein. Let’s call it “gene A.” If the researcher finds another part of the organism’s genome that looks incredibly similar to “gene A” but with a few modifications that make it impossible for the organism to turn it into a protein, that part of the organism’s genome is called a pseudogene.
Since pseudogenes cannot be turned into proteins, it has long been thought that they are the result of a gene being duplicated at some point in history and then being mutated to the point where the gene cannot be used anymore. Indeed, as a commentary in the same issue of Nature says:
Pseudogenes are considered to be defunct relatives of known genes. 2
What Poliseno and her colleagues have conclusively demonstrated is that at least some pseudogenes are anything but defunct, and they might not even be relatives of known genes.
The study focuses on a gene in the human genome called PTEN. This gene produces a protein that is known to suppress tumors. It has been known for a while that there is a region of human DNA that looks a lot like PTEN but does not produce a protein. Thus, that section was identified as a pseudogene of PTEN.3 In their study, Poliseno and her colleagues refer to this pseudogene as PTENP1.
Now remember what evolutionists think is the relationship between PTEN and PTENP1. They think that sometime in the distant past (either in humans or in some ancestor organism), only PTEN existed. Then, it got copied. The copy started to mutate, and it eventually became nonfunctional. At that point, it became the pseudogene PTENP1. That pseudogene is just a leftover vestige of our evolutionary history, and it serves no purpose. As a result, it is a small part of the vast “junk DNA” that makes up (in evolutionists’ minds) the majority of the human genome. In my opinion, Poliseno and her colleagues demonstrate that this evolutionary story is probably not correct.
To understand what this important study demonstrates, you need to learn a couple of things about genetics. First, you need to understand what a gene is and how it instructs the cell to make a protein. A gene is a stretch of DNA that essentially has a recipe for making a specific protein. The stretch of DNA is copied (actually a negative image is made) by a similar molecule called messenger RNA (abbreviated as mRNA). That mRNA then leaves the nucleus (where most of the cell’s DNA is found) and takes the copied recipe to a ribosome, where the protein is made according to the recipe found on the mRNA. Schematically, we could say:
Gene on DNA –> mRNA that leaves the nucleus –> protein
So this is how the PTEN protein is made. The PTEN gene in the DNA produces a messenger RNA (mRNA) that leaves the nucleus and goes to a ribosome, which then makes the PTEN protein that suppresses tumors.
The second thing you need to understand is relatively new to genetics – the existence of microRNAs. In 1993, Lee and his collegues discovered that a roundworm produces short stretches of RNA from its genome. 4 They found this odd, because proteins require long, long stretches of RNA. They didn’t know what possible use could be made of such a small stretch of RNA. As time went on, more and more of these small stretches of RNA have been found, and they have since been called “microRNAs.”
What do microRNAs do? They bind to a small section on mRNA and keep it from being made into a protein. In other words, they regulate the expression of a gene. So…think back to the PTEN gene. The PTEN gene makes a mRNA. The mRNA leaves the nucleus, but on its way to the ribosome, a microRNA might bind to it. If that happens, the PTEN protein won’t be produced.
Now don’t get lost in all this terminology, because it is very important. Let me review this with a figure:
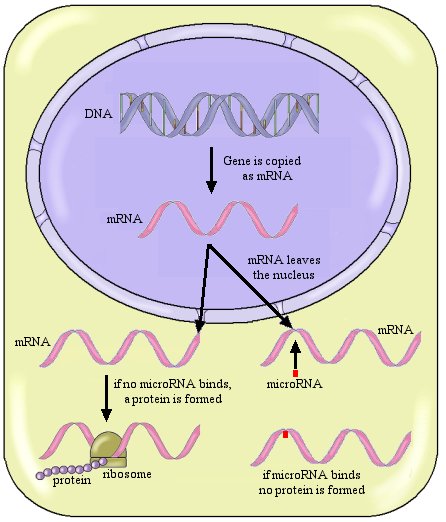
So the general scheme is that once the gene has been copied and turned into mRNA, that mRNA will go to the ribosome and make a protein, unless a microRNA binds to it. If a microRNA binds to the mRNA, no protein is made.
That’s how microRNAs regulate gene expression. They suppress gene expression by stopping the mRNA before it can be used to make the protein for which the gene codes. Geneticists have come to learn that this is a major way in which genes are regulated in nature.
Now you are finally ready to learn what Poliseno and her colleagues found. They found that the mRNA formed by the pseudogene traps microRNAs so that they do not stop the expression of the gene.! In other words, PTENP1 produces decoys that the microRNAs bind to. That way, the microRNAs won’t bind to the PTEN mRNA, and the PTEN protein will be made.
Think about it this way: MicroRNAs suppress the formation of the PTEN protein. So if the cell needs more of the PTEN protein, it can produce decoys using the PTENP1 pseudogene. Those decoys will trap the microRNAs that would have stopped the PTEN protein from forming. As a result, PTEN production will be increased. Thus, while microRNAs suppress gene expression, pseudogenes enhance gene expression!
Poliseno and her colleagues show that this relationship exists not only for this particular gene and its pseudogene, but it exists for other cancer-related genes and their pseudogenes. However, I think that their work reaches far beyond cancer research. As the authors state:
An important implication of our findings is that the decoy mechanism may not be limited to pseudogenes, but may include other long non-coding RNA transcripts including ribosomal RNAs, large intergenic noncoding RNAs (lincRNAs) and coding gene mRNAs5
Indeed, their work indicates that much of what was once considered “junk DNA” is probably not junk at all. Instead, it is a very important part of the genome.
Of course, all of this is exactly what creationists and those in the intelligent design movement have been saying for quite some time now. More importantly, however, it shows once again that supposedly “definitive” evidence for evolution is just not correct. Pseudogenes have been cited as conclusive proof of common ancestry among certain organisms. Indeed, a commenter on this blog cited the similarities between the human and chimp GULO pseudogene as definitive evidence that humans and chimps share a common ancestor. After all, he reasoned, we “know” that the GULO pseudogene is just a defunct, mutated version of the original gene. We see mostly the same random mutations in the chimp GULO pesudogene as we see in the human one, indicating that we both received that pseudogene from a common ancestor.
Of course, this new study shows that pseudogenes probably didn’t even form that way. It was always assumed by evolutionists that a pseudogene is a defunct version of an active gene, because it looks so much like the active gene. What else could possibly explain their similarity? Well, now we know what else. The fact that a pseudogene needs to regulate the expression of the active gene is the reason they are so similar! There is no need to postulate the “mutated copy” explanation for the existence of pseudogenes. There is a simpler (and probably better) explanation: Pseudogenes and genes are both necessary for the production of many proteins. The gene makes the protein, microRNAs reduce the expression of the gene, and pseudogenes increase the expression of the gene. Thus, pseudogenes did not come from genes. They came with genes as part of an exquisitely-designed mechanism for regulated protein production!
REFERENCES
1. Laura Poliseno, Leonardo Salmena, Jiangwen Zhang, Brett Carver, William J. Haveman, and Pier Paolo Pandolfi, “A coding-independent function of gene and pseudogene mRNAs regulates tumour biology,” Nature 465:1033-1038, 2010 full article available online
Return to Text
2. Isidore Rigoutsos and Frank Furnari, “Decoy for microRNAs,” Nature 465:1016-1017, 2010 full article available online
Return to Text
3. Teng DH, et al., “MMAC1/PTEN mutations in primary tumor specimens and tumor cell lines,” Cancer Research 57:5221-5225, 1997
Return to Text
4. Lee RC, Feinbaum RL, Ambros V, “The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14,” Cell 75:843-854, 1993
Return to Text
5. Laura Poliseno, Leonardo Salmena, Jiangwen Zhang, Brett Carver, William J. Haveman, and Pier Paolo Pandolfi, Ibid, p. 1035 full article available online
Return to Text

It’s certainly an elaborate design, but if the cell can control what comes out of the nucleus (as it would have to do for this “junk” to be an effective moderator) why not just do the simple thing and vary the amount of the actual gene as a decoy? Or for that matter why not dump the microRNA and ONLY vary the amount of the gene. I’m sure I’m missing something, but it does look rather over complicated.
Josiah, there are still a lot of things we don’t know about the process of protein synthesis. For example, we don’t really know how the cell “decides” to unwind a portion of the DNA and copy it to begin with. Thus, all I can do is conjecture. However, I can imagine that since it costs energy to unwind DNA, the cell is designed to simply copy everything in an unwound section, without regard to whether or not it is needed. That may be more energy efficient than just unwinding and copying each time something is needed. Of course, that means there might be excess of some proteins made, so the microRNAs (which cost very little energy to make) are made to “stop” the process when the protein isn’t needed. At the same time, just stopping the production of microRNA won’t get rid of those microRNAs that are in the cytoplasm, waiting to pounce. Thus, if the cell suddenly needs the protein, those have to be “cleaned up” by the pseudogene in addition to the cell stopping the further production of microRNAs. Once again, that’s all conjecture, but it makes some sense, at least to me.
I agree that the whole process seems overly complicated, but I expect that a jet engine seems overly complicated to someone who doesn’t understand much about it. I expect that as we learn more about this process, we will find that it results in very efficient protein production regulation. That’s just a guess, though. All we can say for sure right now is vast sections of DNA that evolutionists have dismissed as “junk” have been demonstrated to have a very important function.
BTW, I think your description of the process (elaborate) is better than mine (exquisite).
Putting myself in a darwinist’s shoes for a moment: Wouldn’t darwinian evolution predict this type of scenario? After all, they believe PTENP1 to be a recently (in the darwinian timescale anyway) gene duplication of PTEN. While PTENP1 might no longer code a protein due to the mutation in the start codon it still is a semi-functioning PTEN gene since it hasn’t mutated far enough away yet. To them the ‘function’ of PTENP1 is really just a left-over vestige of the PTEN gene, right? Or am I missing something? I admit I tried to read the paper myself and was having a hard time digesting it all.
Hi Roger. Thanks for the comment! Certainly the “decoy effect” is easy to understand in a Darwinian scenario. As you say, the gene could be copied, mutations at the beginning could make it impossible for the mRNA to produce a protein, but the rest could be similar enough to still trap microRNAs. The problem is that under the Darwinian scenario, that kind of mutation should not be selected, because it would disrupt the cell’s regulation. Remember, the microRNAs are there to reduce the expression of the gene. If copy and mutation were the cause of the pseudogene, then suddenly, there would be a lot of microRNAs trapped by the pseudogene, and as a result, too much expression of the gene. Such a situation should not be selected, as the cell would be less fit to survive given that its regulation system was not working properly.
The other problem is that this is not just one gene/pseudogene pair. The researchers demonstrate the same relationship among other genes and pseudogenes in cancer-related issues, and they postulate that it is a general trend. Thus, you now have to say that this kind of copy and mutation happens a lot, and in each case, the mutations are “lucky” enough to make the gene nonfunctional from the standpoint of protein production but functional from the standpoint of microRNA trapping. Furthermore, the cell somehow has a way of adjusting its gene regulation to account for this new effect each time those lucky mutations occur.
Thanks for the explanation, that makes a lot of sense. This does seem to pose a problem for the darwinian “gene duplication” theory of trait development then (if I’m understanding everything correctly). Duplicated genes it seems then would initially almost always cause a reduction in fitness because:
1. Initially the duplicated gene will cause an overabundance of the coded protein because it is being doubly manufactured (assuming the regulatory elements are also duplicated)
2. If disabled by a nonsense mutation the duplicated gene would still cause an overabundance of the previously coded protein due to microRNA trapping.
Of course, this doesn’t address missense mutations – but it still seems there would be some conflict since both duplicated genes would likely still trap the same microRNA?
I don’t know enough about the underlying chemistry to know if a indel causing a frameshift would greatly eliminate/modify binding sites for microRNA.
Glad to help, Roger. There are many proteins for which too much protein isn’t necessarily a bad thing. Thus, I don’t think #1 applies to all genes. It does apply to tumor-suppressing proteins, as they suppress tumors by reducing the activity of things like reproduction. When a tumor is an issue, that’s a good thing. When a tumor is not an issue, that’s a bad thing. Your #2 is correct, but once again, an overabundance of some proteins isn’t necessarily bad.
Missense mutations would mostly likely still allow for microRNA trapping, unless the mutation happened in the microRNA binding site. Since that is a small part of the gene, however, it isn’t as likely as missense mutations that would keep the microRNA binding site clean.
A frameshift mutation could eliminate the microRNA trapping, depending on where it happens and how much shift occurs. The microRNA is sensitive to where on the mRNA the binding site is. If it moves too far from where it is supposed to be, the microRNA will not bind to it anymore.