
So why is it surprising that the authors of this study found DNA in sediments that are supposed to be “only” 1.4 million years old? There are at least two reasons. First, wet sediment should accelerate the decay of biological molecules (including DNA) compared to the inside of a bone. Thus, DNA should become undetectable much earlier in wet sediment. Second, their results indicate that there would still be DNA found in much “older” sediments, because the “older” the sediment in their study, the more the DNA seem to resist decay.
But wait a minute. This is wet sediment. All manner of microorganisms can infiltrate wet sediment. How do we know that this DNA has really been in the sediment since it was formed? Couldn’t the DNA they detected be from microorganisms that recently started living there? That’s one of the clever aspects of this experiment. The authors looked at DNA that is found in the chloroplasts of cells that do photosynthesis! Since such organisms require light to survive, they wouldn’t have any reason to infiltrate the dark sediments. Even if they did get into those sediments for some reason, they would die in the upper layers, so the deeper (and therefore older) sediments definitely would not harbor any recent microorganisms that had such DNA.
In the end, it seems to me that they really did extract from the sediment DNA that had been there since that sediment had formed. That’s interesting enough, but there is something even more fascinating about what the authors found.
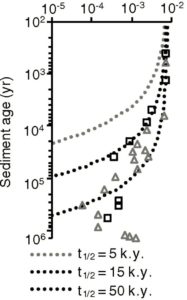 The graph on the left is a slightly modified version of the authors’ Figure 3A. It shows the fraction of chloroplast DNA found versus the assumed age of the sediment. The squares represent what was found in sediment from one site, while the triangles represent what was found in sediment from another site. The various dotted curves show what one would expect if the DNA was decaying the way a biological compound typically decays. They each use a different half-life (5,000 years, 15,000 years, or 50,000 years), but they allow you to compare the data to what is expected based on our knowledge of how DNA decays. Notice that the data don’t follow any of the curves. The “older” the sediment, the “slower” the DNA seems to decay. The authors note this:
The graph on the left is a slightly modified version of the authors’ Figure 3A. It shows the fraction of chloroplast DNA found versus the assumed age of the sediment. The squares represent what was found in sediment from one site, while the triangles represent what was found in sediment from another site. The various dotted curves show what one would expect if the DNA was decaying the way a biological compound typically decays. They each use a different half-life (5,000 years, 15,000 years, or 50,000 years), but they allow you to compare the data to what is expected based on our knowledge of how DNA decays. Notice that the data don’t follow any of the curves. The “older” the sediment, the “slower” the DNA seems to decay. The authors note this:
As described in the Results section, for both sites, the relationship of cpDNA sequence fractions to age best fits a power law (Fig. 3B). This suggests that residual cpDNA is more resistant to degradation in older sediment.
(NOTE: “cpDNA” is their abbreviation for the DNA found in chloroplasts)
The authors suggest that perhaps the older the sediment, the lower the activity from bacteria and other dark-loving microorganisms. That might decrease the rate of DNA decay. Also, they suggest that perhaps some of the DNA is better protected than the rest. For example, some algae (the diatoms) form cell walls made from silica. Theoretically, their DNA should be more resistant to decay than the DNA found in algae that don’t make such cell walls. So maybe the DNA is not decaying as expected because there are several different rates of DNA decay in the sediment, and the only thing they can see is the mixture of all those rates.
I find both of those possible explanations unsatisfactory. The first one is almost certainly not right, because deep sediments support a wide variety of microbial life. The second one probably isn’t right, either. The authors specifically say that chloroplast DNA from diatoms is present in the deeper layers. At some point, you should get to sediment that is old enough that the poorly-protected DNA has decayed away and mostly diatom DNA is left. At that point, the data should follow one of the curves. However, the data never end up following the curves, even in the very “old” sediment.
There is another way to explain what the authors see. If the age of the sediment is calculated wrongly, you wouldn’t expect the data to follow the curves. To me, that’s the more likely explanation. However, more studies will need to be done to figure this out. For example, it would be interesting to see the same kind of graph made exclusively for chloroplast DNA from diatoms. This should give us results for only the better-protected DNA. If my interpretation of the data is correct, you still wouldn’t see anything like the expected decay pattern.
I hope that the authors (as well as others) continue this very interesting line of inquiry.

Dr. Wile, where in the 2012 Allentoft paper on moa DNA do you find that no meaningful DNA can be extracted after 1.5 million years (http://rspb.royalsocietypublishing.org/content/279/1748/4724)? I’m looking at their Table 1, and I don’t see 1.5 million years in there. Is it somewhere else?
In table 1, they have predictions of half-life at different temperatures. For -5 degrees C, they have a half-life of 158,000 years. After 10 half-lives, the DNA would be so degraded that no meaningful sequences would be left. That’s about 1.5 million years.
Allentoft et al. state, “Still, the results indicate that under the right conditions of preservation, short fragments of DNA should be retrievable from very old bone (e.g. greater than 1 Myr). However, even under the best preservation conditions at 258C, our model predicts that no intact bonds (average length = 1 bp) will remain in the DNA ‘strand’ after 6.8 Myr.”
Is the figure they give in Table 1 of 6.8 million years till average length of 1 bp of any significance? I’m trying to make a chart of ancient DNA findings vs. geologic periods, and I’d like to have a reference line indicating the maximum predicted DNA survival time. So, for instance, if it’s 1.5 million years, I’d draw a line there, and any fossils supposed to be older than that should not have detectable DNA. I’m just wondering why you would consider 10 half lives to be the “magic number,” so to speak, as opposed to their 6.8 million year figure.
The 10 half-lives number is a general rule of thumb for biological molecules. Molecules like proteins, DNA, etc. have information in them. As the bonds break, that information gets randomized. Usually, 10 half-lives of decay randomizes the information so much that you can’t see anything meaningful anymore. In a protein, for example, all the molecules’ active sites would have so many breaks in them that they would be utterly useless. For DNA, the sequences would be so small that they wouldn’t contain any useful information.
The Nature article I linked in the OP uses that same rule of thumb:
So 6.8 million years is when the only bases that exist aren’t connected to any other bases. However, for DNA to have meaningful information, several bases have to be connected in sequence. The “10 half-lives” rule of thumb estimates when the sequences become too short to be meaningful.
Dr. Wile, On a slightly related subject, I just got “The Genius of Ancient Man – Evolutions Nightmare”. I wonder if you have read any of Don Landis’ work. The book is pretty simple, but it’s nice to see a Christian tackle these ideas at all. The decreasing of knowledge and complete starting over in almost every civilization is a thorn in the evolutionary framework. The new age crowd has been connecting these dots for quite some time, but they always end up with some fantastic explanation (usually invoking outer space, but sometimes long cycles of time). I’d love to see you write an article on this if you have interest / time to.
Thank You – John
I have read some articles of his online, but I haven’t read any of his books. I am not very knowledgeable about ancient people and their practices, so I am not sure it is something I would write about. However, I have run across a couple of things that I have investigated, like the fact that no one of any significance ever believed in a flat earth and there is a reasonable argument to be made that we are less intelligent than ancient people were. These certainly point to the fact that ancient people were a lot smarter than they are typically given credit for.
You don’t have to go back very far to see the extent of knowledge loss. We only need to go back each decade for the last 20 decades and you will see what we can no longer do. I have various reference materials (engineering, mathematics, programming, etc) that describe various technologies that were “superseded” and forgotten about. Yet, you will see some of these today being redeveloped as “new” ideas.
There are various material scientists, nanotechnology engineers, engineers, and other amateur researchers who are working on projects to discover how people used to do things we cannot do today. Examples include pouring building blocks that are indistinguishable from natural stone or how to lay down uniform thickness thin film layers on ornate structures. The first example is about 2000+ year old technology and the second is 400-600 years old.
There is much knowledge we have lost over the years and that is just in the last 50 years. What did man know that we no longer have access to over the longer term?
I enjoy a set of SF stories from the 40-50’s. In it a specific technology is mentioned. The interesting thing is that the specific technology was a real working technology of the times and I only found out the details of that technology in the last few weeks. Knowing about it now makes more sense within the story structure. The point is that it was a real technology that has since been bypassed by other technologies and has now been forgotten in the main.
I agree.
I would like to bring up another item for Dr. Wile to comment on whenever possible. Hugh Ross from Reasons to Believe and Debora Haarsma from BioLogos had a You Tube discussion that I will try to paste a link for below. A little after the 34 minute mark of the video Hugh Ross makes the comment that ravens far surpass the great apes in intellectual capacity. Given the brain size of birds vs. the brain size of primates, I thought that was a very interesting statement. Dr Ross goes on to explain why he thinks his statement is true. I would be interested in any thoughts Dr. Wile would have on the subject.
https://www.youtube.com/watch?v=FCaGvnmg5_E
I think it is really hard to measure intelligence in animals, but there certainly are indications that Dr. Ross is correct on that point. This study, for example, concludes that crows are more intelligent than apes. On certain mathematical tests, pigeons can even beat human beings. They also have amazing pattern recognition abilities. I do think that a large brain-to-body ratio is not a good indicator of intelligence.
Thanks for your reply Dr Wile and for the links. I am not sure I agree with Dr Ross that Ravens far surpass the great apes in intelligence, but I do think there is good evidence they may have similar or possibly slightly more intelligence when it comes to solving complex problems. The question in my mind was why a creature with a much smaller brain would have similar intelligence to one with a much larger brain. My initial thought was that there may be a higher neuronal density in the Raven than in primates but I had not been able to confirm that suspicion when I emailed you the link for comment.
While I was waiting for your reply I continued to research the subject on line and found a very interesting article that seemed well written and researched. I just recently found the article and read it quickly but didn’t initially realize there was supplemental information available (there were two separate PDF’s). I will paste the link to the article with the supplemental information so you have a chance to read it while I also look at it in more depth.
The article did seem to confirm that there was a greater neuronal density especially in the higher brain areas of birds. This still would not have equaled the number of neurons in higher primates so the authors proposed another possible mechanism to account for greater intelligence in a bird with more densely packed neurons. They suggested that the shorter interneuronal distance that would result from increased neuronal density could result in high speed information processing. I thought that was a reasonable possibility and one that I had not considered until reading their article.
http://www.pnas.org/content/113/26/7255.full.pdf?with-ds=yes
I am not sure you can correlate intelligence to neuron count, neuron density, distance, etc. I would think it would have to do with the network of connections between the neurons. I’ll read the article and see how they make their argument.
Dear Dr. Wile,
I want to notice that in a recent paper, Schweitzer et al. suggested that Allentoft has extrapolated his data too much. They say:
“A recent paper by Allentoft et al. (2012) hypothesizes a half-life for DNA of ~ 521 years in an optimal depositional environment, suggesting that DNA should be degraded to single bases by a little under 7 million years, even though they also state that “considerable sample-to-sample variance in DNA preservation could not be accounted for by geologic age”. Their half-life estimate was based upon extrapolations of data taken from > 150 relatively recent Holocene bones (less than 10,000 years old). Fossils older than this were not examined for DNA. (…) Therefore we suggest more rigorous testing of extrapolation models on actual fossil material from older specimens. Additionally we have found that chemical reactions interfere with analyses of DNA and proteins and should be taken into account before assigning a definitive half-life.”
Source: http://www.sciencedirect.com/science/article/pii/S875632821201318X
Schweitzer, Mary Higby, et al. “Molecular analyses of dinosaur osteocytes support the presence of endogenous molecules.” Bone 52.1 (2013): 414-423.
It’s quite possible that the half-life estimated by Allentoft et. al. is wrong. However, so far, it is the best number we have. Also, we know that the sediment studied here would be worse for DNA preservation than dry bone. Finally, the big deal to me is the fact that the DNA seems more resistant to decay the older the sediment. That doesn’t make any sense if the age of the sediment is being calculated properly.
I wrote on that Schweitzer paper some time ago: DNA and bone cells found in dinosaur bone, as did Dr Wile: Remains of Cells: In DINOSAUR Bones!. It’s rather the opposite of what she claims. Allentoft’s analysis showed, given well attested chemical kinetics, that even if the DNA was frozen at -5°C, it would be totally fragmented in under 7 million years. The DNA was clearly intact enough for an intercalation test like DAPI and propidium iodide to test positive, which would require enough bases for a helical structure. And everyone agrees that dinos flourished in warm climates where the breakdown rate is far higher (normal Arrhenius exponential dependence of rate on temp).
Note also, the discovery of DNA instability was the basis of last year’s Nobel Prize for Chemistry. This lead to realizing that living creatures must have repair machines to cope with the constant chemical onslaught on our heriditary molecule. See DNA repair mechanisms ‘shout’ creation.
Thanks, those are both great articles. The flat earth is a weird one – especially since it’s cropping up so heavily right now (in conspiracy and christian circles). If anything, it illustrates how little trust the populace has in those with the keys to knowledge (government, space agency, private institutions, etc.)
See A flat earth, and other nonsense: Dealing with ideas that would not exist were it not for the Internet.
As a physical chemist, I cringed at the title of the Nature summary paper “DNA has a 521-year half-life”. It’s meaningless to state a half-life of a chemical reaction without stating the temperature, because the rate is exponentially dependent on temp, unlike radioactive decay half lives (at least for the most part, perhaps). This is no reflection on the original authors of course, who did discuss the temperature. Even the Nature paper later does this to some extent, but the headline is still misleading.