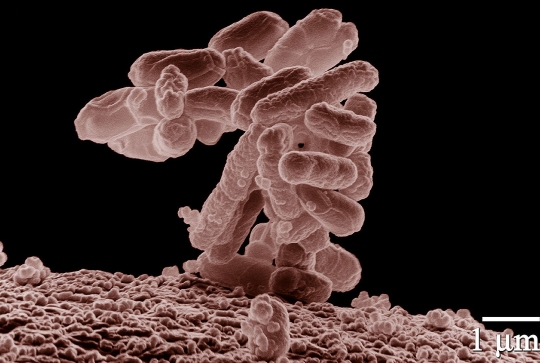
Dr. Richard Lenski, an evolutionary biologist at Michigan State University, has been running a long-term experiment on evolution. Indeed, it has been named the LTEE (Long-Term Evolution Experiment). It started back in 1988 and is still running today. It has followed 12 populations of the bacterium Escherichia coli through more than 50,000 generations, examining how environmental stress changes the bacteria’s genetic and physiological characteristics. More than 6 years ago, I discussed how the project was confirming the creationist view of the genome, and it continues to do just that. In addition, it has inspired another experiment that specifically confirmed a creationist prediction while, at the same time, falsifying an evolutionary one.
To understand what has happened, we need to go back to 2008. In that year, the LTEE showed that even though Escherichia coli normally can’t make use of a chemical called citrate when oxygen is present, one of the their populations developed that ability after 31,500 generations of existence.1 As a result, it was dubbed the “citrate plus” population. How did that happen? At the time, no one knew. However, evolutionists thought it was the result of some rare event or combination of events, exactly the kind upon which evolution depends. New Scientist put it this way:
By this time, Lenski calculated, enough bacterial cells had lived and died that all simple mutations must already have occurred several times over.
That meant the “citrate-plus” trait must have been something special – either it was a single mutation of an unusually improbable sort, a rare chromosome inversion, say, or else gaining the ability to use citrate required the accumulation of several mutations in sequence.
Lenski himself was bold enough to write:
So the bacteria in this simple flask-world have split into two lineages that coexist by exploiting their common environment in different ways. And one of the lineages makes its living by doing something brand-new, something that its ancestor could not do.
That sounds a lot like the origin of species to me. What do you think?
Not surprisingly, a recent experiment has shown that the evolutionary predictions of Lenski and New Scientist are wrong. At the same time, it demonstrated that the predictions of both intelligent design advocates and creationists were correct.
What was the experiment? It was an attempt to reproduce the LTEE’s results – repeatedly. If one of the evolution-inspired predictions was correct, it should be very, very hard to evolve a population of citrate-using Escherichia coli in an oxygen environment. After all, Lenski believed that this was the origin of a new species, and that should be very rare. Indeed, as New Scientist indicated, the ability to use citrate in the presence of oxygen was “something special” that required “rare” conditions. However, there were scientists with another view of the situation. Intelligent design advocates suggested it as well, but I will quote a creationist (Dr. Georgia Purdom) who suggested it:
Mutations which lead to adaptation, termed adaptive mutations, can readily fit within a creation model where adaptive mechanisms are a designed feature of bacteria allowing them to survive in a fallen world. Since E. coli already possess the ability to transport and utilize citrate under certain conditions, it is conceivable that they could adapt and gain the ability to utilize citrate under broader conditions. This does not require the addition of new genetic information or functional systems… (emphasis mine)
If Dr. Purdom is correct, it shouldn’t be at all difficult to produce citrate-using Escherichia coli in an oxygen environment. Adaptive mutations are part of a designed mechanism that produces predictable results in the presence of environmental stress. Thus, given the correct environmental stress, you should repeatedly get the same adaptive result. What did the experiment find? It repeatedly got the Escherichia coli to develop the ability to use citrate in the presence of oxygen, as long as the environmental stress was consistent. Here is how the authors put it:2
Using similar media, 46 independent citrate-utilizing mutants were isolated in as few as 12 to 100 generations. Genomic DNA sequencing revealed an amplification of the citT and dctA loci and DNA rearrangements to capture a promoter to express CitT, aerobically. These are members of the same class of mutations identified by the LTEE. We conclude that the rarity of the LTEE mutant was an artifact of the experimental conditions and not a unique evolutionary event. No new genetic information (novel gene function) evolved. (emphasis mine)
In other words, they were able to repeatedly get the same results as the LTEE very quickly. That means there was no “rare” set of events leading to the ability to use citrate in the presence of oxygen, and this was definitely not any kind of speciation event. Instead, the same genetic changes seen in the LTEE were achieved repeatedly after a short amount of time. This tells us that the ability to use citrate in the presence of oxygen is the result of adaptive mutation, as predicted by Dr. Purdom nearly 8 years ago.
I would also point out the conclusion that I highlighted with boldface type. The researchers conclude that no new genetic information was needed to produce this change. This had been determined back in 2012, confirming what both intelligent design advocates and creationists had predicted, but it is nice to see it clearly spelled out in the scientific literature.
REFERENCES
1. Zachary D. Blount, Christina Z. Borland, and Richard E. Lenski, “Historical Contingency and the Evolution of a Key Innovation in an Experimental Population of Escherichia coli,” Proceeding of the National Academy of Sciences, 105:7899–7906, 2008
Return to Text
2. Van Hofwegen DJ, Hovde CJ, and Minnich SA, “Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA,” Journal of Bacteriology 198:1022-34, 2016
Return to Text

Hello Dr, I came here to post a good blog writen by an ID professor in Brazil(yes I’m brazilian) http://pos-darwinista.blogspot.com.br
Thank you for the link, Victor! I can’t read the Spanish portions, but the English portions are interesting.
Doctor, May you answer me why the amoeba has such a giant DNA base? Please.
I don’t think anyone knows the answer to that question, Victor. All we do know is that the size of the genome doesn’t seem to correlate with the complexity of the organism. The Japanese canopy plant has a huge genome as well. So does the marbled lungfish. Since we know very little about DNA, we have no idea how to interpret the size of the genome. Hopefully, as we learn more about DNA, we will learn how the length of a genome affects the way the organism uses it.
I’ve heard that some of the previous amoeba DNA estimates were too large by an order of magnitude. For example see Taft et al, 2007:
“…amoebae are often cited as the most dramatic example of the lack of correlation between genome size and biological complexity. There are may problems with this conclusion, including a likely variation in ploidy… as well as the presence of significant amounts of contaminating DNA from their prey. The amoeba genome is probably smaller than 20 pg, far less than the 700 pg commonly cited” http://salamander.uky.edu/srvoss/425SP08/Taftetal.pdf
However 20pg is still almost 7 times larger than a human genome. If it helps, I wrote an article about Junk DNA and have a section in it with some ideas about variations in genome sizes. http://notascientist.d512.com/worldview/biology/evolution/junk-dna/#Objection-The-C-Value-Paradox
The article you wrote is excellent, Joe. It contained several things of which I was unaware. Thank you for linking it!
My guess is that the capability to use citrate in the presence of oxygen is an ability that was coded in the bacteria’s DNA all along. Environmental conditions trigger the organism to mutate in such a way that it turns on that functionality as needed. It’s not creating information, but rather just setting a parameter.
Consider the eyeless fish. When fish dwell for generations in the absence of light, a parameter is set that causes the fish to stop growing eyes, so that the fish doesn’t expend energy maintaining an organ that will be useless to it. That’s the same process as with the bacteria, only in reverse, setting a parameter to turn off a function.
I highly suspect that these bacteria that gained the ability to use citrate in the presence of oxygen could also lose that ability again in just a few generations, just like they have found that fish can lose their eyesight “in the merest handful of generations”.
http://www.biosciencetechnology.com/articles/2014/01/rapid-evolution-method-found-eyeless-fish
In the article to which I linked, scientists who refuse to consider Intelligent Design call this parameter-setting type of mutation “rapid evolution”, a term I find hilarious!
I remember reading somewhere that researchers had taken a species of blind fish or salamander and gotten them in a few generations to produce eyes again. Anybody know about that?
This may be the study you are thinking about, David:
Restoring Sight in Blind Cavefish
They made hybrids from different species of blind cavefish, assuming that each species went blind along a different adaptive path. As a result, if you started hybridizing the fish, you would get working genes from one species that weren’t working in the other, and vice-versa, resulting in working eyes. That idea seemed to be confirmed by the experiment.
Response to JoeCoder’s reply on May 23 – wow. I just now got around to reading your posted article, and I’m glad I did.
Your data compression analogies certainly ring true with me. And the performance optimization. It seems like genomes may have something going on like what we would call “loop-unrolling optimization”, a time-space tradeoff well know to us coders. https://en.wikipedia.org/wiki/Loop_unrolling