When a scientist refuses to see the design that is so obvious in nature, it can lead to all sorts of incorrect conclusions. Consider, for example, transposable elements in DNA. Often called “transposons,” they jump around in an organism’s genome. In other words, they are in different places in different cells of the same organism. Those who have their naturalist blinders on initially thought that they were useless – part of the “junk DNA” that represents all the evolutionary “flotsam and jetsam” that has accumulated over hundreds of millions of years. Dr. Leslie Pray, writing in Nature Education, puts it this way:
Transposable elements (TEs), also known as “jumping genes” or transposons, are sequences of DNA that move (or jump) from one location in the genome to another. Maize geneticist Barbara McClintock discovered TEs in the 1940s, and for decades thereafter, most scientists dismissed transposons as useless or “junk” DNA. McClintock, however, was among the first researchers to suggest that these mysterious mobile elements of the genome might play some kind of regulatory role, determining which genes are turned on and when this activation takes place.
Of course, we now know that these supposedly useless stretches of DNA have widespread functionality throughout the genome. However, a recent study demonstrated that one set of transposable elements (the HERV-H subfamily) has a particularly interesting function, which indicates that a creation scientist’s prediction I wrote about nine years ago has been confirmed.
Let’s start with what the HERV-H subfamily is. The “ERV” part stands for “endogenous retrovirus.” When a retrovirus infects a cell, its RNA is converted into DNA, and that DNA is then inserted in the cell’s genome. That way, the retrovirus is replicated as a part of the cell’s DNA. As long as the cell stays alive, that stretch of DNA stays in the genome. If the cell reproduces, each of its progeny also has that DNA sequence. So these stretches of DNA are thought to be the result of retrovirus infections that happened in our evolutionary past. The first “H” stands for human, because this particular group of DNA sequences is found in humans. The last “H” is a category that separates this HERV family from other HERV families.
You can see why most scientists dismissed ERVs as useless. They came from an infection in our primate ancestors, so we have no use for them. I distinctly remember my university biology professor specifically indicating that HERVs are excellent evidence for evolution. ERVs are “obviously” useless, and yet we have many in common with apes. While similar stretches of DNA that have purpose can be understood in terms of common design as well as common ancestry, similar stretches of DNA that are useless are best explained by common ancestry. Of course, now that we know how functional HERVs are, they can also be explained by common design.
But that’s not the point of this post. In the study I am writing about today, the authors determine one of the things the HERV-H subfamily does: It helps to change the way the DNA is arranged in the nucleus of the cell, which allows each type of cell to be as efficient as possible. Remember, every cell has the same DNA (barring some mutation or chimeric effect). However, once a cell is mature, it has a specific job to do, like being a skin cell. To do its job efficiently, it must use its DNA differently than another type of cell, like a nerve cell. As a result, the DNA in each cell’s nucleus is arranged to maximize its efficiency.
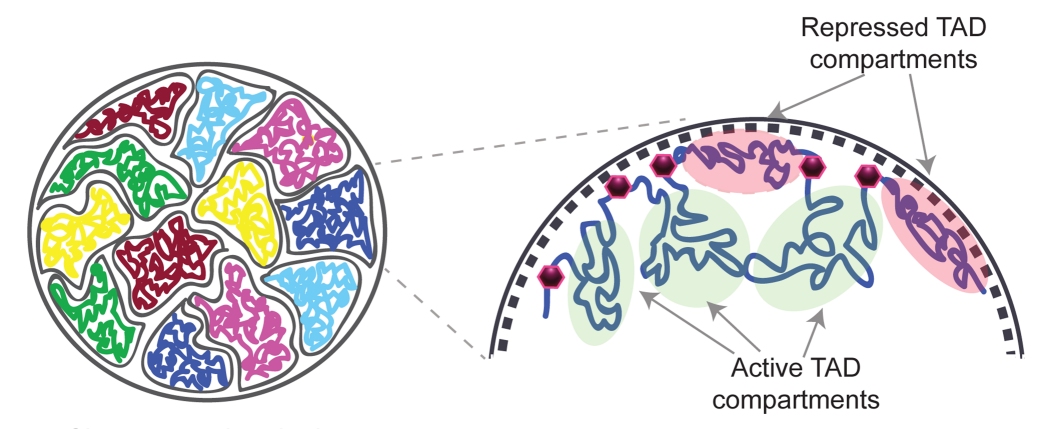
You can see that by looking at the image above. Each chromosome has its own “territory” in the nucleus. Each colored region on the left represents a chromosome’s territory. Within each territory, the DNA of the chromosome is not spread out evenly. There are areas called topologically associating domains (TADs) that have to interact quite a bit. Those areas are placed so that they can interact easily, while the areas that don’t need to interact much are “pushed away” near the territory’s edge. Each type of cell will have its own arrangement of TADs, since each type of cell uses its DNA differently.
How does the cell know how to arrange its TADs? That’s where the study comes in. The authors show that the HERV-H subfamily is at least partially responsible for TAD arrangement.
Here, by interrogating chromatin reorganization during human pluripotent stem cell (hPSC) differentiation, we discover a role for the primate-specific endogenous retrotransposon human endogenous retrovirus subfamily H (HERV-H) in creating topologically associating domains (TADs) in hPSCs.
Now remember, human pluripotent stem cells are cells that have the ability to become many different kinds of cells. So as a stem cell matures into whatever cell it is going to be, the HERV-H subfamily helps to arrange the DNA in the nucleus so the mature cell can do its job efficiently.
How does this confirm a creationist prediction? In April of 2009, Dr. Peer Terbor wrote about ERVs. In the article, he develops a hypothesis for the origin of certain viruses. To develop that hypothesis, he specifically redefined ERVs as “Variation Inducing Genetic Elements” (VIGEs): parts of the genome that are designed to change the DNA when change is needed. What is the HERV-H subfamily doing? It is inducing variation in the DNA. Now Dr. Terbor was specifically talking about the variation you see from one generation to the next, but implicit in his definition is that HERVs have a specific function: they produce changes in the DNA that help the organism to survive. That’s what the HERV-H subfamily is doing, so his prediction has been confirmed.
If you have heard the lie that creationism cannot make useful predictions about the way the world works (a necessary component of any scientific theory), you should know that this is just one of many confirmed creationist predictions. You can read about others here.

That is radical. Seems like HERVs are like dog ears on a page in a recipe book. Does this explain all HERVs? Do you think they are actually from viruses or originally designed pieces of the genome? Why did we come to think they are remnants of retroviruses in the first place?
Also, I’m sure you saw the new discovery about DNA and the H bonds holding the strands together have been “disproved”. What’s your take?
I don’t think this explains all HERVs, and most likely, even HERV-Hs do other things. However, I do think all HERVs are functional and have nothing to do with previous infections. They are a designed part of the genome. People came to the conclusion that they are retroviral remnants because of the blinders of evolution. They see that the elements behave like retroviruses, they assume evolution is true, so they conclude that these must be remnants from infections in our ancestors. I think Terborg’s speculation is more likely to be true. They behave like retroviruses because retroviruses started as ervs and ended up escaping the genome.
I plan to write a blog post about the DNA study. I haven’t waded through the paper yet.
Is it correct to say that these HERVs are then, at least partially, responsible for beginning and maintaining cell differentiation?
I would say they are at least partially responsible for producing cell differentiation. We don’t know that they begin the process, and once a cell has differentiated, it doesn’t need help maintaining the differentiation, at least as far as I know. Without the HERVs, however, it appears that cells would not fully differentiate.