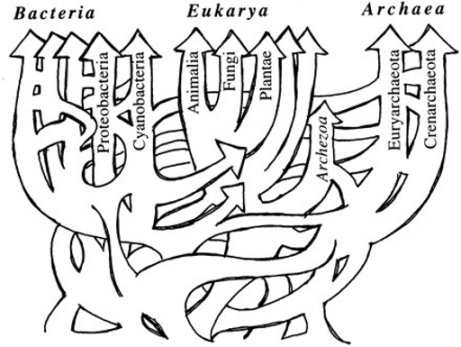
When I was a young, impressionable student at university, I was taught as fact that all organisms on this planet could be arranged in a hypothetical “tree of life” that showed how all of them evolved from a single, common ancestor. It didn’t matter that no such tree had actually been constructed. I was told that in time, we would be able to sequence DNA quickly and efficiently, and once that happened, the tree of life would emerge from the data in all its glory. However, once DNA analysis did become reasonably quick and efficient, the tree of life never emerged. Instead, the supposed evolutionary relationships that had been determined from the fossil record were contradicted by those that were determined from the genes. Worse yet, the genetic story of evolution changed depending on which specific genes were studied.
This was especially apparent in the analysis of single-celled organisms. As more and more genetic analyses were done on such organisms, it became increasingly obvious that there was simply no way to arrange their genetic information into any pattern that even remotely resembled a hypothetical tree of life. Some scientists understood what this meant: there is something seriously wrong with the evolutionary framework to which biologists have been clinging. As a result, they have started investigating other, more promising paradigms such as creationism or intelligent design. However, most biologists continue to cling to a view that has been falsified again and again by various data. As a result, they had to find a “way out.” They had to find some means by which they could explain around the fact that the genetic data falsified the tree of life.
Enter the concept of Horizontal Gene Transfer, also known as lateral gene transfer. In this process, a section of DNA is transferred from the genome of one organism to the genome of another, unrelated organism. In other words, rather than passing down a gene through the process of reproduction, horizontal gene transfer allows a gene to travel horizontally between unrelated organisms.
Now the phenomenon itself was known long before the problem with the tree of life had been documented. Way back in 1960, for example, Japanese scientists determined that an antibiotic-resistant bacterium could transfer its resistance to an unrelated bacterium that was not resistant to the antibiotic.1 In time, the mechanism was fully worked out, and it was demonstrated that bacteria could, indeed, swap genes back and forth. As it became increasingly clear that this was a common phenomenon among bacteria, it was recognized that horizontal gene transfer could “smear out” the tree of life for single-celled organisms.2
Since horizontal gene transfer was so successful at explaining away the failed evolutionary prediction of the tree of life when it came to single-celled organisms, it’s not surprising that this same “way out” is now being used to explain why certain genes in animals do not fit the pattern predicted by the evolutionary hypothesis.
In a study published recently in the Proceedings of the National Academy of Sciences of the United States of America, investigators were studying a section of DNA called “BovB.”3 This DNA segment is called a retrotransposon. Although retrotransposons were long thought by evolutionists to be useless junk, studies have demonstrated that they are important regulators of how genes are expressed.4 Well, this particular retrotransposon is found in all sorts of animals, such as cattle, horses, platypuses, geckos, snakes, sea urchins, zebrafish, etc. The investigators decided to compare the BovB found in animals to see if they could find some sort of pattern.
Well, they found a pattern, but it wasn’t what the hypothesis of evolution predicted. In evolutionary terms, you would expect the BovB found in these different animals to vary according to their supposed evolutionary relationships. Two animals that are closely related in evolutionary terms should have similar BovB, while those that are distantly related should have more variation in their BovB. Instead, several exceptions to that expectation were found. For example, the cattle’s BovB was more similar to that of the snake than that of the horse, and the BovB of the gecko was more similar to that of the horse than to that of other reptiles.
How did the authors deal with this problem? They invoked horizontal gene transfer. They suggested that BovB must have been transferred from one species of animal to another at least nine different times. How did this horizontal gene transfer happen? The authors aren’t sure, but they did find two species of ticks that feed on reptile blood and have the BovB retrotransposon in their genome. They suggest that perhaps blood-sucking ticks are responsible for the horizontal gene transfer.
I find this study to be very interesting, but I am surprised by how the authors characterize their results:
Our results provide compelling evidence for HT of genetic material that has transformed vertebrate genomes. [Note: HT abbreviates “horizontal transfer”]
In my view, the evidence is only compelling if you believe that all these animals must be related by evolution. After all, the study only showed that BovB does not conform to evolutionary expectations. Rather than being evidence for a hypothesis, then, it is really evidence against a hypothesis. In the end, horizontal gene transfer is simply invoked to explain around the fact that the data do not fit the proposed evolutionary pattern. Thus, this is only evidence for horizontal gene transfer in animals assuming that the proposed evolutionary relationships must be 100% correct!
Now don’t get me wrong. I am not claiming that horizontal gene transfer didn’t happen in this case. It might very well have. Then again, it might not have; it might be that the proposed evolutionary relationships are just completely wrong. I find that to be a more likely explanation, but I could very well be incorrect. The point is that until we at least find a mechanism by which this horizontal gene transfer took place, it’s far too soon to talk about “compelling evidence!”
REFERENCES
1. Akiba, T.; Koyama, K.; Ishiki, Y.; Kimura, S.; and Fukushima, T., Japanese Journal of Microbiology 4(2):219-27, 1960
Return to Text
2. W. Ford Doolittle and Eric Bapteste, “Pattern pluralism and the Tree of Life hypothesis,” Proceedings of the National Academy of Sciences of the United States of America, doi:10.1073/pnas.0610699104, 2007
Return to Text
3. Ali Morton Walsha, R. Daniel Kortschaka, Michael G. Gardner, Terry Bertozzia, and David L. Adelsona, “Widespread horizontal transfer of retrotransposons,” Proceedings of the National Academy of Sciences of the United States of America, doi:10.1073/pnas.1205856110, 2012
Return to Text
4. Wenhu Pia, Xingguo Zhua, Min Wua, Yongchao Wanga, Sadanand Fulzeleb, Ali Erogluc, Jianhua Linga, and Dorothy Tuana, “Long-range function of an intergenic retrotransposon,” Proceedings of the National Academy of Sciences of the United States of America, doi:10.1073/pnas.1004139107, 2010
Return to Text

There is nothing that evolution can’t do! The more improbable the results, the more powerful it must be. The more we falsify it, the more amazing it is.
Unfortunately, Greg, there seem to be a lot of people who think that way!
Have there been any observed instances of horizontal gene transfer in larger creatures, or has this only been seen with singled celled organisms? If it can only happen with single celled oragnisms, that would sure make the evolutionary tree of life look pretty odd! It would be shaped more like round bush than a tree, as all other life forms branched out from a few single celled ancestors that swapped their genes around.
Keith, I don’t know of any observed examples of horizontal gene transfer in animals. However, there have been some inferred ones. For example, sequencing the genomes of yeast and another fungus showed regions that look very bacteria-like. It is thought that they are the result of horizontal transfer from bacteria to the fungi. Also, a bean beetle that has a mutualistic relationship with a bacterium seems to have some of that bacterium’s genes in its genome, suggesting horizontal gene transfer. These strongly suggest horizontal gene transfer from bacteria to animals, but the actual transfer has not been observed. By contrast, horizontal gene transfer has been directly observed between single-celled organisms.
I see. So scientists have seen things that look like the results of HT in larger animals, but not the process itself. You’re right; it’s definitely a little early to be talking about “compelling evidence.”