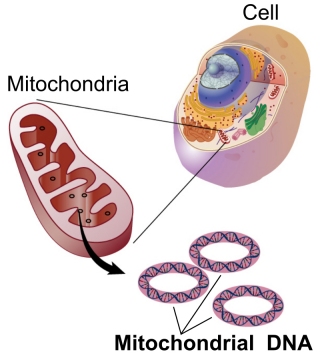
The authors of the study analyzed the DNA of many different species of animals. However, they did not look at the DNA found in the nucleus of the cell. That DNA, called nuclear DNA, is responsible for most of an organism’s genetically-defined traits. They looked at mitochondrial DNA, which is the small amount of DNA that is found in the mitochondrion, the structure that produces most of the energy that the cell ends up using. To give you an idea of how different mitochondrial DNA is from nuclear DNA, the nuclear DNA of a human being is over 3 billion base pairs long, while human mitochondrial DNA is just over 16,000 base pairs long. You don’t need to know what “base pairs” are to see that there is only a tiny, tiny amount of mitochondrial DNA in a human cell compared to nuclear DNA.
Now even though there isn’t a lot of mitochondrial DNA, some sections of it seem to be very characteristic of the species of animal from which the cell comes. For example, a 2016 study analyzed a section of mitochondrial DNA (called the COX1 gene) among different species of birds. It showed that the COX1 gene alone was enough to separate 94% of the birds into species. Similar studies indicate that the COX1 gene can separate other species of animals, so the sequence of the COX1 gene is often referred to as the DNA barcode of the animal. This is what the authors of the study I am discussing focused on.
The authors analyzed the “DNA barcodes” of many different animal species, looking for a specific kind of mutation: one that doesn’t actually change the product of the gene. Remember, a gene is a “recipe” that codes for a particular protein. The COX1 gene used in the DNA barcode is the recipe for a protein lovingly called “mitochondrially encoded cytochrome c oxidase I.” A protein is a string of amino acids linked together, and the DNA “recipe” is just a list of those amino acids and the order in which they need to be strung together. Well, the genetic code has more than one way of indicating an amino acid, and sometimes, a mutation will change part of the DNA, but the resulting change will indicate the same amino acid. This is called a synonymous substitution, because the mutation doesn’t change the recipe for the protein. It just changes the way that recipe reads.
Consider, for example, a recipe for a cake. It might call for two teaspoons of vanilla extract. Suppose you copy that recipe so you can give it to a friend, but instead of writing “two teaspoons of vanilla extract,” you write “2 tsp. vanilla extract.” You have essentially made two synonymous substitutions, substituting a number for the word “two” and an abbreviation for the word “teaspoon.” A synonymous substitution mutation in DNA is similar. The product of the recipe doesn’t change, but the way the recipe reads changes. As a result, even after the mutation happens, the protein produced by DNA remains the same.
The authors assume that synonymous substitution mutations can happen freely, because they don’t change the proteins being produced and therefore should have no effect on the organism. Because of that, synonymous substitution mutations should not be “weeded out” by natural selection, so the more synonymous substitution mutations, the longer those mutations have been accumulating. When they compared the number of these mutations among many different animals, they found that 90% of them had the same level of mutations, indicating that they had been experiencing mutations for the same length of time. This is consistent with the idea that 90% of the animals came into being at the same time.
Now, of course, the authors don’t argue for that idea! It would invalidate the evolutionary hypothesis, and I am sure they don’t want to be considered heretics! As a result, they try to explain around their conclusion by suggesting that there are mechanisms which tend to reduce the genetic diversity among individuals in a species. Because of these mechanisms, most species had uniformity in their mitochondrial DNA at roughly the same time and have just been collecting mutations since then.
As I have said, I am skeptical of their conclusion. That’s because of their assumption that synonymous substitution mutations don’t affect the organism and are thus not subject to natural selection. They produce some arguments to support that assumption, but they discount the most important argument against their assumption: the fact that we know synonymous substitutions change the rate at which a gene is turned into a protein. Since the mitochondrion is the “powerhouse” of the cell, it would make sense to me that its function is highly dependent on the rate at which its genes can be turned into proteins. As a result, I find it highly unlikely that the organism is unaffected by synonymous substitution mutations.
Now while I disagree with the more “sensational” conclusion of their study, they do make one statement that I think is quite important:
The tight clustering of barcodes within species and unfilled sequence space among them are key facts of animal life that evolutionary theory must explain.
In other words, they are saying that there is a lot of discontinuity in mitochondrial DNA of animals. You have a sequence that defines one species, a sequence that defines another species, and nothing in between. If you hope to come up with any kind of plausible mechanism for evolution, you will need to explain that kind of discontinuity.

Hi, doesn’t evolution posit that most animal species came into existence at the same time since they all ‘evolved’ from a ‘common ancestor’ and so each would have the same amount of time to evolve? In Darwinian thought, all organisms are the ‘end of the root’ of the ‘tree of life,’ and so *all* of them have lineages that came into existence at the same time. Or… perhaps this study is suggesting that the ‘divergences’ had occurred at the same time???
Your last sentence is correct. The study is looking at how long the mitochondrial genome of each species has been accumulating mutations. Thus, it is trying to see when each species diverged. Yes, evolution thinks all animals are related through a common ancestor, but each species arose slowly over time. Jawless fish, for example, were supposed to have evolved 510 million years ago, while cartilaginous fish (like sharks) and bony fish (like tuna) were supposed to have evolved about 90 million years later. Mammals weren’t supposed to have evolved until about 200 million years after that. Thus, the fish should have had a lot more time to accumulate mutations than the mammals. If this study is really sensitive to how long the species have existed, then the fish should show a lot more mutations than the mammals.
It’s funny to know that the cosmic panspermia theory was dying. But, for my surprise, a new study says otherwise. Might be useful to read it:https://www.sciencedirect.com/science/article/pii/S0079610718300798
God Enlighten you all!
I’m having a bit of trouble understand some parts of the paper. The on-line article*says;
And yet—another unexpected finding from the study—species have very clear genetic boundaries, and there’s nothing much in between.
“If individuals are stars, then species are galaxies,” said Thaler. “They are compact clusters in the vastness of empty sequence space.”
Does this mean that all species in a genera are clustered together and isolated from others? Or just individuals of the same species?
*https://m.phys.org/news/2018-05-gene-survey-reveals-facets-evolution.html
According to the study, it’s the species that are isolated from one another in their DNA barcodes. Obviously, the species in a genus are clustered more closely than two species from two different genera. However, they are saying that the species are distinct enough in their DNA barcode that they are isolated from the other species.
“In other words, they are saying that there is a lot of discontinuity in mitochondrial DNA of animals. You have a sequence that defines one species, a sequence that defines another species, and nothing in between”
Is this something that Baraminologists could use (or are using) to define “Kinds” ? The NCSE website broadly states “Baraminologists also utilize DNA/DNA hybridization and blood serum reactivity measurements to determine baraminological divisions.” – This seemed a little vague to me.
I don’t think this would be helpful in determining kinds, since a kind would be all animals that are related by a common ancestor. However, I would think that any theory of diversification within kinds would have to be able to explain the discontinuity in the barcodes.
I would love to read your opinion on this video which claims to describe proof for evolution.
I hope you find it interesting and worthwhile.
What is the evidence for evolution – https://www.youtube.com/watch?v=lIEoO5KdPvg
The video is simply wrong. First of all, the “tree of life” as they call it is in complete disarray, because genetic evidence doesn’t agree with fossil evidence, and even when you restrict yourself to just genetics, different genes give different “trees.” More importantly, the focus of the video, cetaceans, do not support evolution. You can see that here, here, here, and here.
If I may, the video might seem convincing… if all you watch is that video. There are major problems with the Theory of Evolution: holes in evolutionary theory, and failed evolutionary predictions. It is easy to make the evidence “look” like it supports evolution by being selective. Notice that the video only focuses on a couple features of whale anatomy.
The transition of land mammal to sea mammal would actually entail many more changes than what is discussed in the video. The video chooses to focus on *just* those morphological features that are easily explained by evolution. However, it does not address the thousands of changes that would be necessary for such a transition: different skin, different respiratory system, different bone structure, etc.
This is a good, realistic critique on “whale evolution” by Dr. David Berlinski:
https://www.youtube.com/watch?v=IZdlQK0cOlI