For those of you who do not speak nerd, “AFK” means “Away from Keyboard.” I will be traveling for the next three weeks, and for much of that time, I will not have internet access. That means I will not be able to moderate or respond to your comments. However, please feel free to leave your comments. When I have internet access, I will try to do what I can to moderate the comments and answer any questions you might have. I will also try to post new articles, but it’s not clear whether or not that will happen. It’s possible that there won’t be a new article posted for as many as three weeks.
Month: January 2013
Being Degenerate Can Be Very Good!
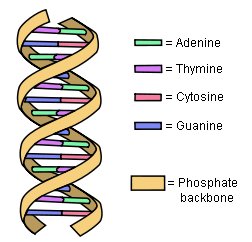
How does a gene list the amino acids? As shown in the illustration above, it does so by using the four nucleotide bases known as cytosine (C), guanine (G), thymine (T), and adenine (A). A group of three nucleotide bases codes for a specific amino acid. For example, when a gene has three thymines in a row (TTT), this means “use the amino acid called lysine.” When it has three guanines in a row (GGG), it means “use the amino acid called proline.” So by grouping its four nucleotide bases three at a time, a gene specifies which amino acid should be used in building a protein.
Here’s the catch: There are only 20 amino acids in the standard proteins of life. As a result, there need to be only 20 codes to specify them. However, there are 64 possible ways you can group four nucleotide bases three at a time. Thus, there are 64 different possibilities for how a gene can specify an amino acid, but there are only 20 amino acids the gene needs to specify. As a result, most amino acids are specified by more than one set of three nucleotide bases. As I said above, a sequence of three thymines (TTT) means “use the amino acid called lysine.” However, two thymines followed by a cytosine (TTC) means the same thing. This is why we say the genetic code is degenerate. It has multiple ways it can specify most amino acids.
Poop Transplants Treat Clostridium difficile Infections!
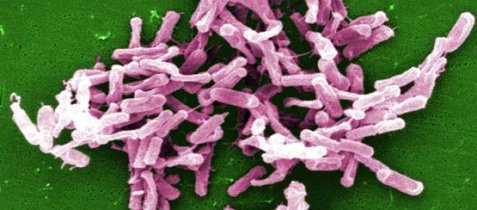
Clostridium difficile is a bacterium that produces toxins which can kill intestinal cells and cause severe inflammation of the intestinal walls. Mild infections from this bacterium can cause diarrhea, while severe infections can cause death. Over time, C. difficile infections have gotten worse. As one medical resource states:1
C. difficile infection has transformed from a nuisance into a potentially life-threatening illness with an attributable mortality rate of up to 16.7%.
The typical treatment for severe cases of C. difficile infection is a round of strong antibiotics, but that doesn’t always work. A significant number of patients end up experiencing one or more recurring infections within 60 days.2 As a result, medical researchers are trying to come up with new ways to treat this infection.
In a recent study published in the New England Journal of Medicine, researchers tested an…interesting…approach. They took 43 patients who had a C. difficile infection and split them into three groups. The first received the standard antibiotic treatment (in this case, 500 milligrams of vancomycin four times per day for 14 days). The second received the standard antibiotic treatment plus a bowl lavage 4 or 5 days into the treatment. In case you aren’t familiar with the term, a bowel lavage involves flushing out the intestines. It is usually done to prepare the intestine for medical imaging. The third group was given a shortened round of antibiotics (500 milligrams of vancomycin four times per day for 4 or 5 days), a bowel lavage, and then a poop transplant.
Yes, you read that right. After the bowel lavage, the patients were given a mixture of water, salt, and the feces from a healthy donor. Now don’t worry. They didn’t have to eat or even smell the stuff. It was sent into their intestine through a sterile tube that went up the nose, down the esophagus, through the stomach, and into the start of the small intestine. While the process sounds incredibly gross, the results were amazing!
Continue reading “Poop Transplants Treat Clostridium difficile Infections!”
Astronomers are Finally Starting to Question an Absurd Assumption
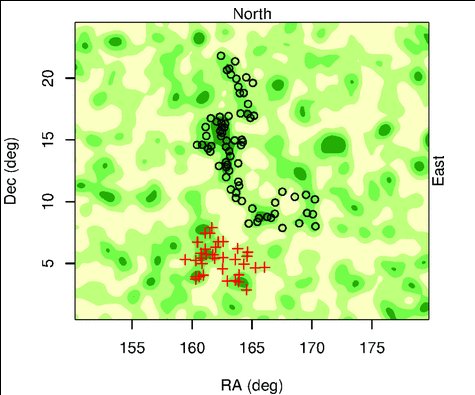
One of the more absurd assumptions that is routinely made in astronomy is called the cosmological principle. One way to phrase the principle is:
Viewed on a large enough scale, the properties of the universe are the same no matter where you are.
However, observations have never supported this assumption. Instead, the observable universe seems incredibly “lumpy,” with huge structures separated by vast areas devoid of structures. Nevertheless, cosmologists have doggedly taken the cosmological principle as their starting assumption when it comes to developing models of the universe, despite the fact that observations don’t support it.
Indeed, the cosmological principle is a necessary starting point for the Big Bang, which most, but certainly not all, astronomers think is a good description of the origin and development of the universe. As Paul Fleisher says in his book, The Big Bang:1
The cosmological principle is the central idea of the Big Bang theory. This rule says the universe is homogeneous and isotropic at very large scales.
Even if we go away from the Big Bang model, the vast majority of models that attempt to describe the universe start with the assumption that the cosmological principle is valid. There are some models that do not start with that assumption, but they are few and far between.2
I have always been skeptical of the cosmological principle, simply because it isn’t supported by observation. The universe doesn’t look homogeneous at all. Instead, it looks really “lumpy.” Nevertheless, when I read the scientific literature, the cosmological principle seems to be considered a fact in almost all of the astronomy-related papers.
It looks like that might be starting to change.
Continue reading “Astronomers are Finally Starting to Question an Absurd Assumption”
Horizontal Gene Transfer: Another “Way Out” for Evolutionists
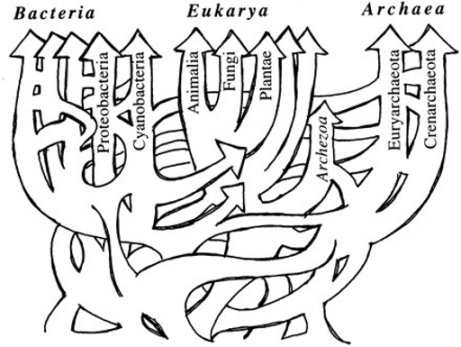
When I was a young, impressionable student at university, I was taught as fact that all organisms on this planet could be arranged in a hypothetical “tree of life” that showed how all of them evolved from a single, common ancestor. It didn’t matter that no such tree had actually been constructed. I was told that in time, we would be able to sequence DNA quickly and efficiently, and once that happened, the tree of life would emerge from the data in all its glory. However, once DNA analysis did become reasonably quick and efficient, the tree of life never emerged. Instead, the supposed evolutionary relationships that had been determined from the fossil record were contradicted by those that were determined from the genes. Worse yet, the genetic story of evolution changed depending on which specific genes were studied.
This was especially apparent in the analysis of single-celled organisms. As more and more genetic analyses were done on such organisms, it became increasingly obvious that there was simply no way to arrange their genetic information into any pattern that even remotely resembled a hypothetical tree of life. Some scientists understood what this meant: there is something seriously wrong with the evolutionary framework to which biologists have been clinging. As a result, they have started investigating other, more promising paradigms such as creationism or intelligent design. However, most biologists continue to cling to a view that has been falsified again and again by various data. As a result, they had to find a “way out.” They had to find some means by which they could explain around the fact that the genetic data falsified the tree of life.
Enter the concept of Horizontal Gene Transfer, also known as lateral gene transfer. In this process, a section of DNA is transferred from the genome of one organism to the genome of another, unrelated organism. In other words, rather than passing down a gene through the process of reproduction, horizontal gene transfer allows a gene to travel horizontally between unrelated organisms.
Now the phenomenon itself was known long before the problem with the tree of life had been documented. Way back in 1960, for example, Japanese scientists determined that an antibiotic-resistant bacterium could transfer its resistance to an unrelated bacterium that was not resistant to the antibiotic.1 In time, the mechanism was fully worked out, and it was demonstrated that bacteria could, indeed, swap genes back and forth. As it became increasingly clear that this was a common phenomenon among bacteria, it was recognized that horizontal gene transfer could “smear out” the tree of life for single-celled organisms.2
Since horizontal gene transfer was so successful at explaining away the failed evolutionary prediction of the tree of life when it came to single-celled organisms, it’s not surprising that this same “way out” is now being used to explain why certain genes in animals do not fit the pattern predicted by the evolutionary hypothesis.
Continue reading “Horizontal Gene Transfer: Another “Way Out” for Evolutionists”
Male DNA in Female Brains? Yes!
(Public domain image)
How in the world did male DNA get into these women’s brains? The researchers aren’t sure, because they don’t have detailed medical histories for most of the women. However, the most likely explanation is that the DNA comes from the male children that these women carried. I have written about this phenomenon, called fetomaternal microchimerism, before. As I mentioned in that article, when I first heard about a baby leaving cellular remains inside his or her mother, I thought it couldn’t possibly be true. However, I was wrong. There is solid evidence to suggest that not only do babies leave a lasting, cellular imprint on their mothers, mothers do the same for their babies.
However, the possibility that children leave some of their DNA behind in their mother’s brain is very surprising. After all, the cells that make up the brain are incredibly sensitive. In fact, the contents of your own blood are toxic to your brain cells. As a result, you have an elaborately designed blood-brain barrier that shields your brain cells from your blood. This barrier is so vigilant that it allows only certain substances (such as the glucose and electrolytes that the brain cells need) to pass through it. As a result, your brain cells are protected from the majority of substances found in your bloodstream.2
Boring Math Drills Aid Higher Math Skills

When I comment on math, I always like to make it clear that I am not, in any way, a mathematician. Indeed, because many homeschooling parents requested it, I actually tried to write a mathematics program. Based on that experience, I can make one very definitive statement: While I use a lot of math in my field, I am definitely not an expert at it. When it comes to mathematics education, you need to take everything I say with a grain of salt.
Nevertheless, one thing I have always stressed when it comes to homeschooling is to emphasize math in the elementary years. Based on my experience with students from junior high all the way through graduate school, I have come to the conclusion that the best way to produce excellent science and mathematics students is to spend a lot of time on math in the elementary years. This not only includes learning new mathematical concepts, but it also means drilling your student in math. In my experience, the best science and math students are the ones who spent a lot of time on math drills when they were young.
An interesting study that seems to support this view was recently published in the Journal of Neuroscience.1
Continue reading “Boring Math Drills Aid Higher Math Skills”
Bacteria in Breast Milk? Yes!
Scientists now call the collection of bacteria that lives in a person’s body the microbiome, and as the article linked above indicates, each person seems to have his or her own special mix of bacteria in that microbiome. Indeed, some researchers think that analyzing the DNA of the bacteria a criminal leaves behind can aid in identifying that criminal in cases where his or her own DNA is not available at the crime scene or too degraded to analyze properly.1
So where does an infant start collecting the bacteria that will make up his or her own microbiome? One of the sources is the breast milk that the infant drinks. It has been known for quite some time that breast milk contains bacteria, but the details have not been well studied. However, a group of Spanish researchers have begun to shed some light on those details. They studied the breast milk of 18 mothers who varied in weight, weight gain during pregnancy, and the mode in which the baby was delivered. They sampled the milk these mothers produced at three different times: the first secretions of milk produced after giving birth (called colostrum), the milk that was produced one month after giving birth, and the milk that was produced six months after giving birth. They sequenced the DNA of the bacteria found in these samples of milk, and they came up with some amazing results.2
More Evidence Against Feathered Dinosaurs
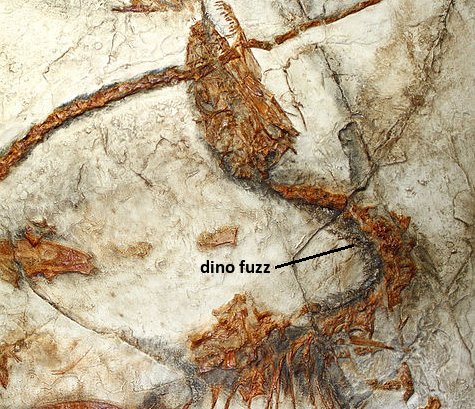

Several months ago I wrote an article about the fossil evidence for primitive feathers (often called “protofeathers”) in some dinosaur specimens. The article discussed a study by Theagarten Lingham-Soliar, Alan Feduccia, and Xiaolin Wang that provided strong evidence against the common interpretation that the dinosaur Sinosauropteryx was covered in such protofeathers. In the discussion that followed, Dr. Jonathan Sarfati suggested that I should read another article by Lingham-Soliar.1 Over the holidays, I finally had a chance to do so. As he suggested, this is another very important study in the “feathered dinosaurs” debate. While Dr. Sarfati has his own excellent analysis of this study and a few others, I would like to add some thoughts of my own.
Lingham-Soliar’s study focused on two fossils: NIGP 127587 (identified as a Sinosauropteryx fossil) and GMV 2124 (thought to be a Sinosauropteryx fossil). Both exhibit exceptional preservation. In fact, the latter fossil is so well-preserved that the stomach contents were analyzed and three mammal skulls were found! Two came from mammals in the genus Zhangheotherium, and the third came from the genus Sinobaatar.2 The important aspect of the fossils for this study, however, is the fact that both show some sort of “fuzz” extending from the body of the animal (pointed out in the picture above). This “fuzz” has been routinely interpreted to be the remains of primitive feathers, but Lingham-Soliar and his colleagues strongly dispute that interpretation.
In an attempt to understand precisely what this “fuzz” represents, Lingham-Soliar performed a detailed examination of the fossils and also did a simple experiment. The combination of his fossil analysis and the results of the experiment provide still more evidence that Sinosauropteryx did not have any feathers.
Continue reading “More Evidence Against Feathered Dinosaurs”
